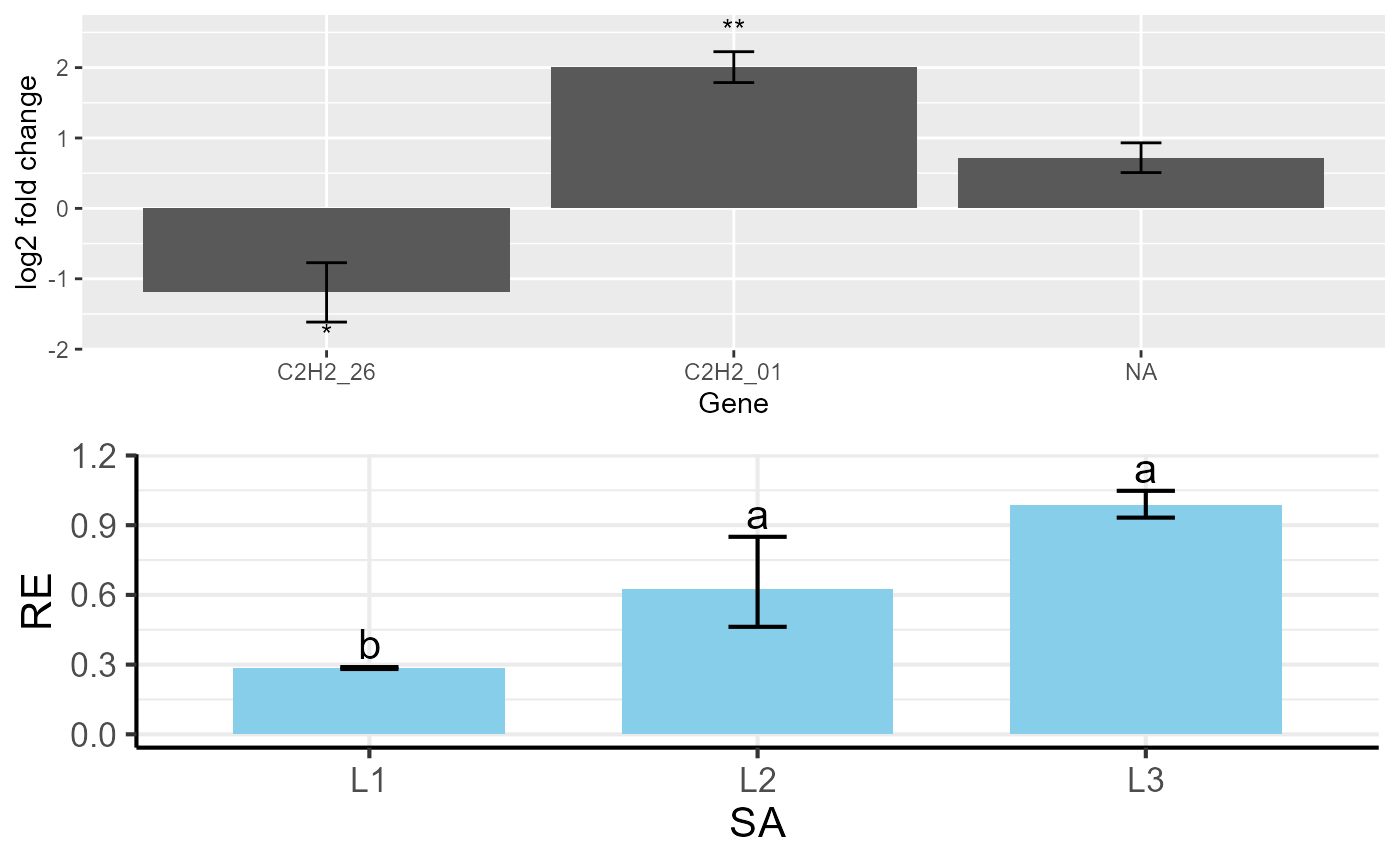
The multiplot function arranges multiple ggplot2 objects
into a single plotting layout with a specified number of columns.
Details
Multiple ggplot2 objects can be provided either as separate
arguments via ....
The function uses the grid package to control the layout.
Author
Pedro J. (adapted from https://gist.github.com/pedroj/ffe89c67282f82c1813d)
Examples
# Example using output from TTEST_DDCt
data1 <- read.csv(system.file("extdata", "data_ttest18genes.csv", package = "rtpcr"))
out <- TTEST_DDCt(
data1,
paired = FALSE,
var.equal = TRUE,
numberOfrefGenes = 1)
#> *** 18 target(s) using 1 reference gene(s) was analysed!
#> *** The Control level was used as calibrator.
p1 <- plotFactor(out,
x_col = "gene",
y_col = "log2FC",
Lower.se_col = "Lower.se.log2FC",
Upper.se_col = "Upper.se.log2FC",
letters_col = "sig")
p2 <- plotFactor(out,
x_col = "gene",
y_col = "RE",
Lower.se_col = "Lower.se.RE",
Upper.se_col = "Upper.se.RE",
letters_col = "sig")
# Example using output from ANOVA_DCt
data2 <- read.csv(system.file("extdata", "data_1factor.csv", package = "rtpcr"))
out2 <- ANOVA_DCt(
data2,
numOfFactors = 1,
numberOfrefGenes = 1,
block = NULL)
#>
#> Relative Expression
#>
#> gene SA dCt RE log2FC LCL UCL se Lower.se.RE
#> 1 PO L1 1.81000 0.28519 -1.81000 0.18404 0.44193 0.02082 0.28111
#> 2 PO L2 0.67333 0.62706 -0.67333 0.40466 0.97167 0.43880 0.46261
#> 3 PO L3 0.01667 0.98851 -0.01667 0.63792 1.53178 0.08413 0.93252
#> Upper.se.RE Lower.se.log2FC Upper.se.log2FC sig
#> 1 0.28934 -1.83082 -1.78918 b
#> 2 0.84996 -1.11213 -0.23453 a
#> 3 1.04787 -0.10080 0.06746 a
#>
#> Note: Using default model for statistical analysis: wDCt ~ SA
df <- out2$relativeExpression
p3 <- plotFactor(
df,
x_col = "SA",
y_col = "RE",
Lower.se_col = "Lower.se.RE",
Upper.se_col = "Upper.se.RE",
letters_col = "sig",
letters_d = 0.1,
col_width = 0.7,
err_width = 0.15,
fill_colors = "skyblue",
alpha = 1,
base_size = 14)
# Combine plots into a single layout
multiplot(p1, p2, cols = 2)
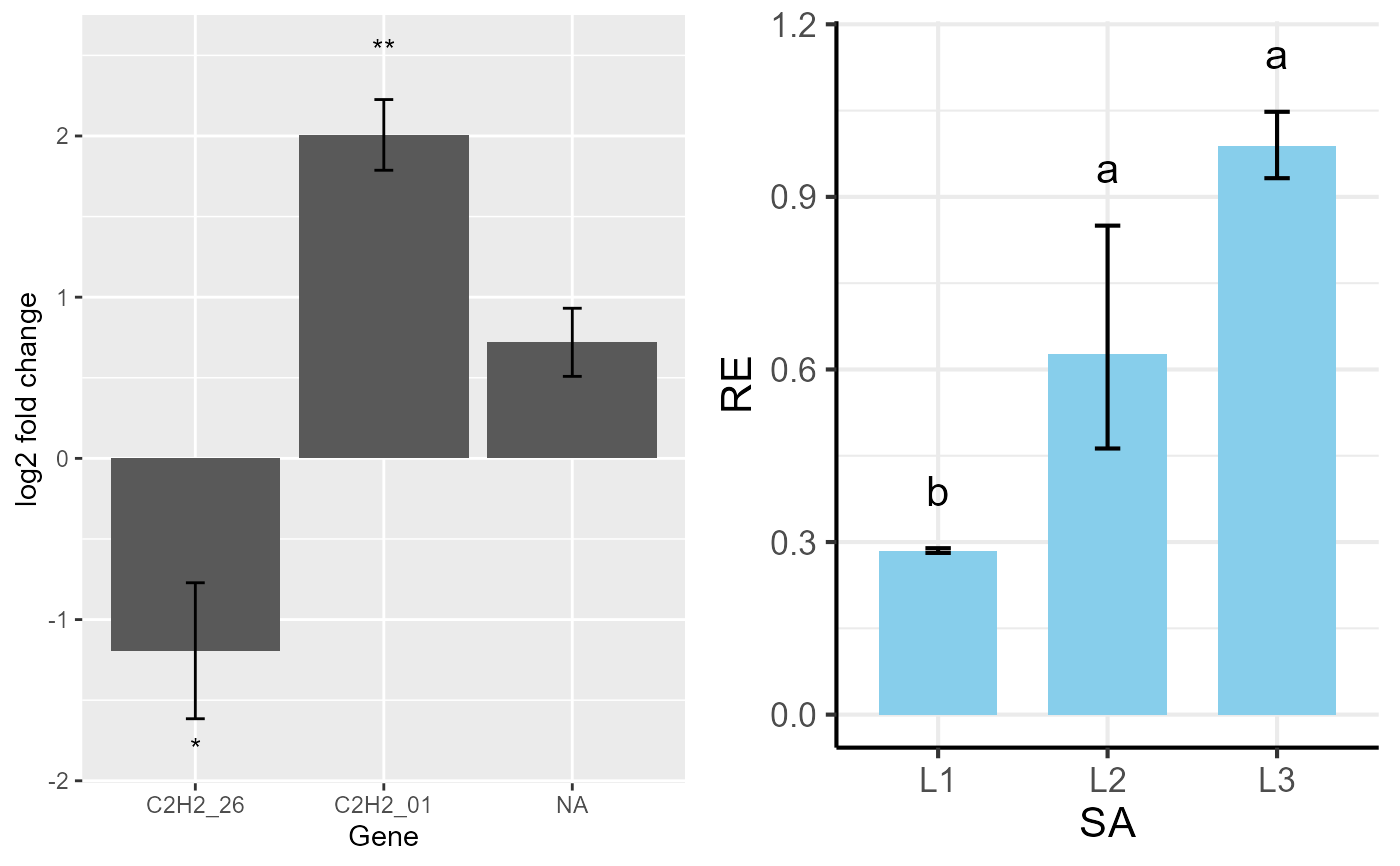 multiplot(p1, p3, cols = 2)
multiplot(p1, p3, cols = 2)